GlycoForm & Glycologue: two software utilities for the display of N-glycans
Description:
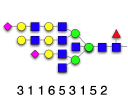
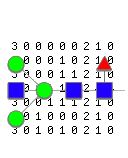
GlycoForm draws N-glycan structures in symbolic or text form, using the 9-digit encoding system devised by Krambeck and Betenbaugh [1].Glycologue accepts lists of such 9-digit codes, displaying the corresponding structures in a grid.
Both applications are written in REALbasic.
Update (November 2014):
A web application edition of GlycoForm is now available, in conjunction with O-Glycologue, a simulator of the enzymes of O-linked glycosylation. These are based on string-based structure identifiers that use single-letter codes to represent common monosaccharides and brackets to represent branch points in complex oligosaccharides.
System Requirements
- Mac OS X 10.4 or higher / Windows 9x or higher / Linux x86
Downloads
| GlycoForm | [Mac OS X] [Windows] [Linux x86 with GTK] | |
| Glycologue | [Mac OS X] [Windows] [Linux x86 with GTK] |
Installation
Download the archive appropriate to your operating system and decompress it to a local drive. The application can be started from any location of your choosing.Important note for Windows users. In addition to the executable file (.exe), the zip archive will contain a folder dynamically loaded libraries (DLLs). This folder (called either "GlycoForm Libs" or "Glycologue Libs") must be present at the same location as the executable. Therefore, it is recommended that you use the "Extract all files" option after double-clicking on the zip file.
Sample Glycologue data are available for download as either a zipped or tar-gzipped archive.
Click here to download the project source code.
References
[1] Krambeck, F.J. and Betenbaugh, M.J. (2005) A mathematical model of N-linked glycosylation. Biotech. Bioeng. 92(6), 711-728.[2] McDonald, A.G., Tipton, K.F., Stroop, C.J.M. and Davey, G.P. (2010) GlycoForm and Glycologue: two software applications for the rapid construction and display of N-glycans from mammalian sources. BMC Res. Notes 3, 173. [PMID: 20565879]